Licensing
Flexible Modular Access Tailored to Your Pipeline
Licensing
Flexible Modular Access Tailored to Your Pipeline
Available as a full enterprise platform or licensed capability modules.
Select a module of interest to explore its capabilities:
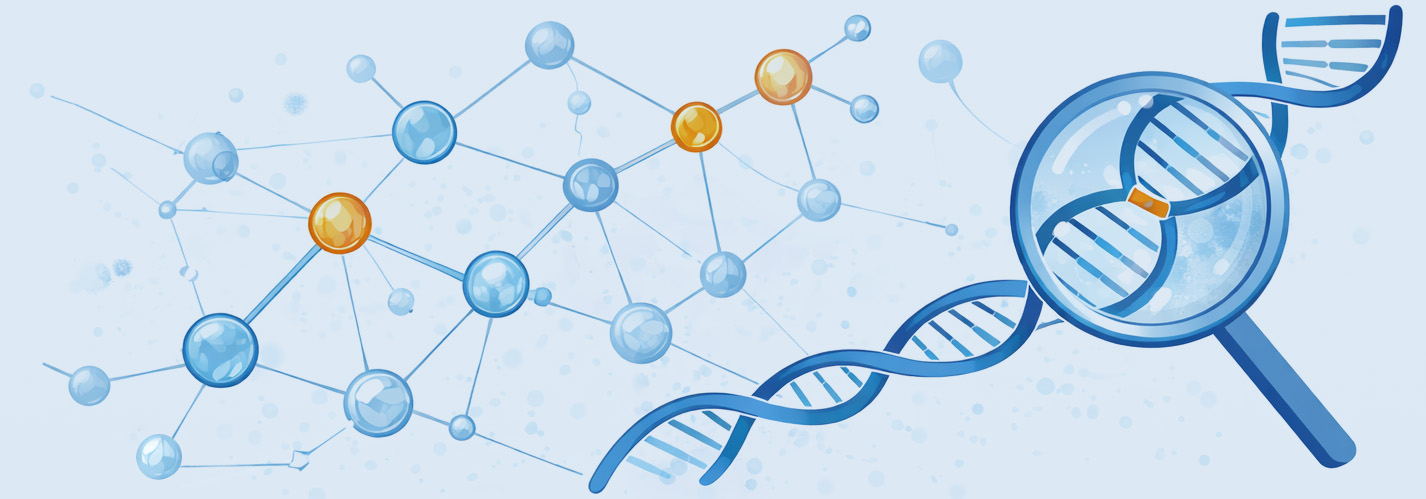
Moves beyond gene associations to establish why a gene drives disease – delivering validated causal graphs, confidence-scored targets, and regulatory-grade mechanistic evidence packages.
Causal Engine Output
- Directional molecule-to-molecule causal relationships – not correlations
- Ranked target list with Tier 1–3 Causal Confidence Scores
- Validated causal graph from genetic variants through to clinical outcomes
- Full evidence dossier with provenance tracking for regulatory submission
- Patient Specificity Index (PSI) safety labelling on every finding

Six Included Workflows
- Causal Discovery — find novel drivers of disease or phenotype
- Directed Causality Testing — validate “does X cause Y?”
- Intervention Ranking — prioritise targets by actionability
- Comparative Causality — compare mechanisms across cohorts
- Counterfactual Analysis — simulate “what if” perturbations
- Evidence Inspection — explain any causal claim in full
Technical Details
Causal DAG Construction
Genetically anchored Directed Acyclic Graph using PC, NOTEARS, and FCI algorithms. Four-factor directional arbitration: genetic anchoring, temporal priority, perturbation power, and cellular context.
Mendelian Randomization
MR-Egger, Inverse Variance Weighted (IVW), and eQTL-based causal effect estimates. Genetic instruments provide randomisation-equivalent causal evidence without confounding.
Temporal Validation
Granger causality analysis and directional temporal tests using pseudotime and time-series data. Rules out reverse causation and establishes time-ordering of causal events.
Causal Confidence Scoring
CCS synthesis across all four validation streams. Tier 1 (Driver), Tier 2 (Strong Candidate), Tier 3 (Candidate) classification with contradiction detection and PSI safety labelling.
Multi-Modal Data Pillars
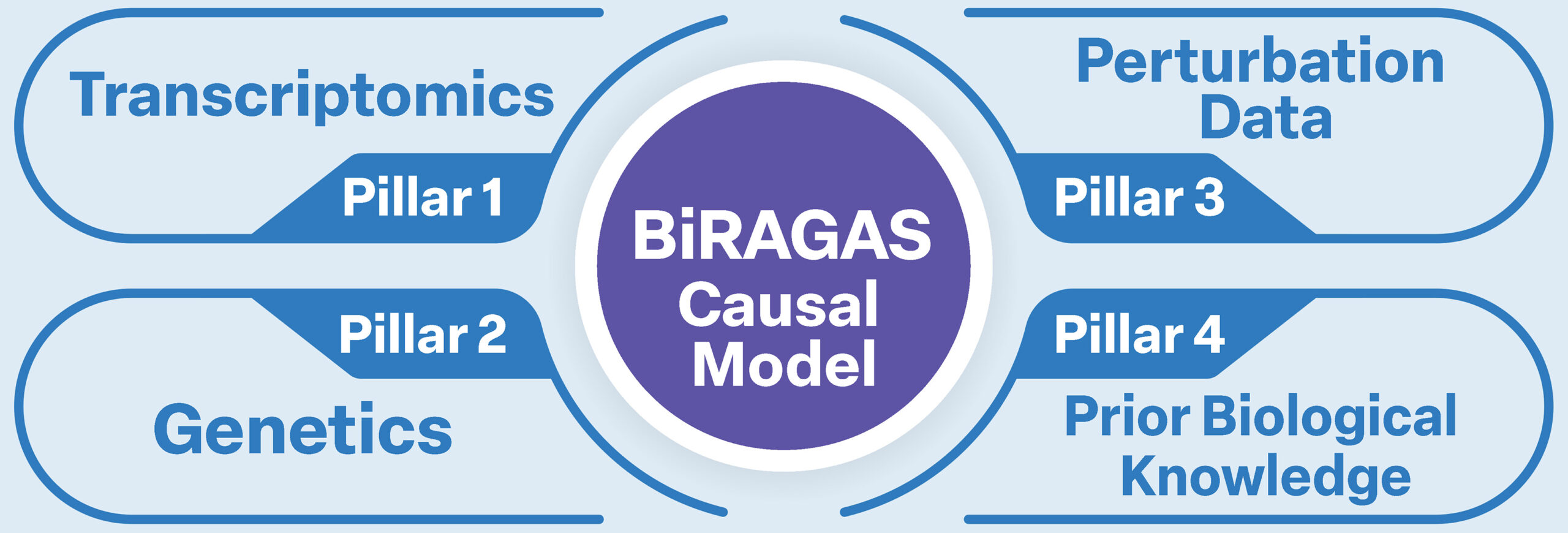

Transcriptomics
Patient cohort expression profiles with DESeq2 differential expression across disease states and conditions.
Genetics
GWAS summary statistics, eQTLs, and MR estimates providing genetically anchored causal evidence.
Perturbation
CRISPR DepMap essentiality and drug EC50 data as interventional validation of causal claims.
Prior Knowledge
SIGNOR, REACTOME, KEGG, and Gene Ontology constraining and contextualising discovered relationships.
Perturbation Intelligence
End-to-end CRISPR analysis across three complementary modules – from large-scale functional screens to precise targeted editing characterisation – providing the interventional evidence base that confirms therapeutic target relevance.
Three integrated modules
- Perturbation Validation – integrate DepMap and CausalBench data to confirm nominated target relevance
- CRISPR Screening – genome-wide and focused-library screen analysis with statistical hit calling
- Targeted CRISPR – amplicon-seq, DNA pooled, and transcriptomic perturbation analysis
CRISPR Suite Output
- Gene essentiality profiles confirming functional relevance of causal candidates
- Ranked essential genes, synthetic lethal pairs, and therapeutic vulnerabilities
- Editing efficiency quantification and on-target indel profiling
- Downstream transcriptomic consequences of targeted edits at gene-level resolution
- Direct integration with Causal Engine intervention ranking and target scoring
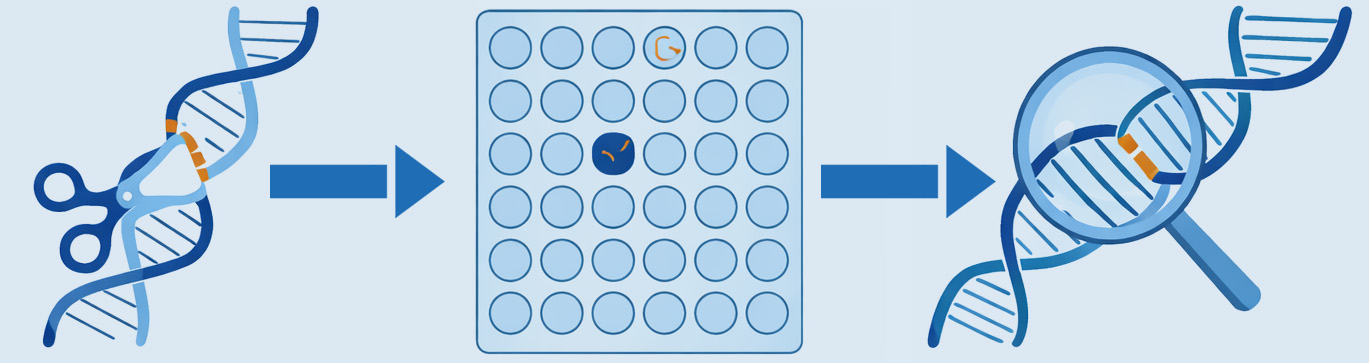
Technical Data
End-to-end CRISPR analysis across three complementary modules – from large-scale functional screens to precise targeted editing characterisation – providing the interventional evidence base that confirms therapeutic target relevance.

Integrates DepMap gene essentiality screens and CausalBench Average Causal Effect (ACE) scores to quantify the phenotypic consequence of each candidate gene knockout. ACE scores directly inform the Causal Engine’s directional arbitration — genes with high ACE are enforced as causal upstream of lower-ACE targets. Drug EC50 data and disease signature reversal scoring complete the validation picture, confirming druggability and directional effect on disease state.
- DepMap essentiality across cancer cell line panels
- ACE scores from CausalBench perturbation experiments
- Drug EC50 dose-response validation
- Signature reversal scoring for disease-state shift assessment
Technical Data

Full analysis pipeline for genome-wide and targeted pooled CRISPR screens. Processes guide RNA counts from raw sequencing output, applies quality control, performs statistical hit calling with FDR control, and generates ranked gene-level essentiality and enrichment results. Supports both positive and negative selection screens, synthetic lethality analysis, and pathway-focused sub-library experiments.
- Genome-wide and focused library screens
- Guide-level QC and normalisation
- Statistical hit calling with FDR control
- Synthetic lethality and combination vulnerability identification

Precise, locus-level characterisation of editing outcomes using three complementary modalities. Amplicon sequencing quantifies editing efficiency and on-target indel profiles at nucleotide resolution. DNA pooled data enables population-level analysis of editing outcomes across conditions. Transcriptomic CRISPR perturbation (CRISPRi/CRISPRa) captures downstream gene expression consequences of targeted edits, resolving biological mechanism at the pathway level.
- Amplicon-seq: editing efficiency and indel profiling
- DNA pooled: population-level editing outcome analysis
- Transcriptomic perturbation: CRISPRi/CRISPRa expression consequences
- End-to-end: edit → transcriptional effect at single-gene resolution
Therapeutic Intelligence
Translates any causally identified gene or target directly into the full therapeutic landscape – drugs, mechanisms, safety signals, pharmacogenomics, and clinical trial evidence – across 8.6 million curated biological vectors.
Drug Agent Output
- Gene-to-drug translation: surface associated drugs, mechanisms, and clinical phase
- Safety profiling: adverse event signals from FDA FAERS with significance scoring
- Precision medicine matching: genotype-specific efficacy, dosing, and toxicity guidance
- Clinical trial context: active and completed trials for every identified target
- Single unified query returning an integrated therapeutic report
Knowledge base at a glance

Technical Details – Knowledge Collection
Type: Genes
Source: GeneALaCart
Primary Use: Gene-level annotation, function, pathway membership, and disease associations for every queried gene
Type: Targets
Source: Open Targets
Primary Use: 9,521 targets · 3,000 diseases · 3,000 drugs · 16,729 associations for foundational therapeutic recommendation
Type: Drugs
Source: Open Targets Enriched
Primary Use: Full mechanism of action, indications, linked targets, diseases, and clinical phase for drug prioritisation
Type: Safety
Source: FDA FAERS
Primary Use: Pharmacovigilance records with log-likelihood ratio significance scores for safety flagging and contraindication alerts
Type: Variants
Source: PharmGKB · ClinPGx
Primary Use: Variant-level efficacy, dosing, and toxicity with rs IDs, genotypes, and clinical evidence levels
Type: Associations
Source: Curated KG
Primary Use: Mechanism of action, route, indication, interactions, adverse reactions, molecular targets, efficacy scores
Type: Compounds
Source: ChEMBL
Primary Use: Chemical compound records for structure–activity relationship context and compound-level biological annotation
Type: Full KG
Source: Structured CSV KG
Primary Use: Drug name · cancer status · mechanism · route · indications · interactions · adverse reactions · targets · efficacy scores
Genomics & Transcriptomics Platform
A comprehensive multi-modal genomics platform spanning data acquisition, harmonisation, expression analysis, cell composition profiling, single-cell analysis, and multi-omics biomarker discovery – the analytical foundation that feeds into BiRAGAS and stands alone as a full genomics research platform.
Four capability pillars
Data Acquisition & Harmonisation
GEO, ArrayExpress, TCGA retrieval with ComBat batch correction
Expression Analysis & FASTQ Pipeline
End-to-end RNA-seq: QC → alignment → DEG → pathway
Cell Type Deconvolution & Single-Cell
xCell, CIBERSORT, Bisque + full scRNA-seq pipeline
Multi-Omics Biomarker Discovery
xCell, CIBERSORT, Bisque + full scRNA-seq pipeline
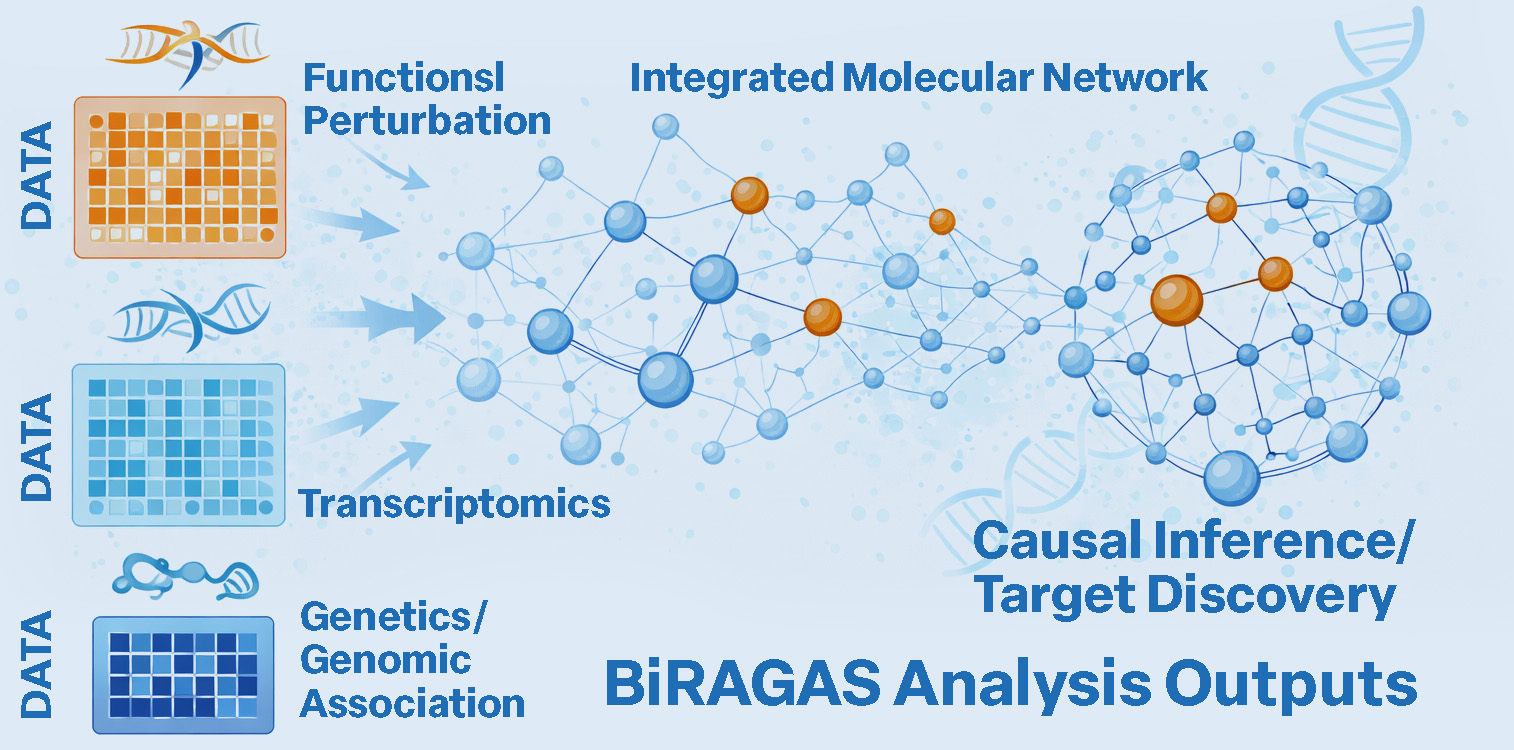
- Harmonised multi-cohort expression datasets ready for causal analysis
- Differential expression and pathway enrichment reports
- Cell-type composition estimates for every bulk RNA-seq sample
- Single-cell clustering, annotation, and trajectory analysis
- Cross-omics biomarker signatures for patient stratification and treatment response
- Outputs formatted for direct ingestion into the Causal Engine
Data Acquisition & Harmonisation
Automated retrieval from NCBI Gene Expression Omnibus (GEO), EMBL-EBI ArrayExpress, and The Cancer Genome Atlas (TCGA). The harmonisation pipeline resolves batch effects and metadata inconsistencies across multi-study datasets using ComBat for microarray and bulk RNA-seq data, ComBat-Seq optimised for count-based sequencing, and additional normalisation strategies (TMM, RLE, quantile, VST) selected by data type and experimental design.
- GEO, ArrayExpress, TCGA automated retrieval
- ComBat and ComBat-Seq batch correction
- TMM, RLE, quantile, and VST normalisation
- Metadata standardisation across multi-cohort studies
Expression Analysis & FASTQ Pipeline
End-to-end RNA-seq processing from raw FASTQ files through count matrices, differential expression, and pathway enrichment. Quality control and adapter trimming (FastQC, Trimmomatic), alignment and quantification (STAR, Salmon), differential expression with DESeq2, and gene set enrichment analysis against KEGG, Reactome, and Gene Ontology databases – fully automated and optimised for downstream causal inference.
- FastQC and Trimmomatic QC and trimming
- STAR and Salmon alignment and quantification
- DESeq2 differential expression with FDR control
- GSEA pathway enrichment against KEGG, Reactome, GO
Cell Type Deconvolution & Single-Cell Analysis
Three validated deconvolution methods estimate cellular composition from bulk RNA-seq: xCell for enrichment scoring across 64 immune and stromal cell types; CIBERSORT for immune fraction estimation using SVR on gene expression signatures; and Bisque for reference-based decomposition using single-cell data as a biological reference. For single-cell studies, the full scRNA-seq pipeline provides preprocessing, normalisation, dimensionality reduction, clustering, cell type annotation, and trajectory analysis.
- xCell — 64 cell type enrichment scoring
- CIBERSORT — immune fraction estimation via SVR
- Bisque — reference-based bulk deconvolution
- Full scRNA-seq pipeline: QC → clustering → annotation → trajectory
Multi-Omics Biomarker Discovery
Integrates genomic, transcriptomic, proteomic, and metabolomic data using MOFA (Multi-Omics Factor Analysis) to extract shared latent factors representing the molecular axes of variation most predictive of disease state or treatment outcome. Cross-platform data harmonisation enables joint embedding across omics layers. Biomarker candidates are ranked by cross-omics consistency and biological interpretability, producing multi-modal signatures for patient stratification, diagnostic development, and treatment response monitoring.
- MOFA — latent factor extraction across omics layers
- Cross-platform harmonisation and joint embedding
- Cross-omics biomarker ranking by consistency and effect size
- Multi-modal disease signatures for stratification and diagnostics
Enterprise Deployment
All five modules can be licensed together as an integrated enterprise deployment – with bespoke configuration, proprietary data integration, custom knowledge base development, and a dedicated scientific partnership team. Enterprise access includes white-label deployment, co-development arrangements, and regulatory submission support.
Proprietary Data Integration
Internal datasets – unpublished cohort data, in-house CRISPR screens, compound libraries – securely integrated into programme-specific knowledge bases isolated from other users.
Regulatory & Publication Support
Full evidence provenance documentation, mechanistic summary reports, and expert scientific review for FDA submission packages and peer-reviewed publication.
Co-Development Options
Strategic partners contributing novel data, methodology, or disease expertise may access co-development and IP co-ownership arrangements.
